ARGONNE LABS: LCHS
Lakes Community High School students have been given the opportunity to work with Argonne Scientist Nathan Guisinger as part of the Exemplary Student Research Program. Matthew Anderson, Andrew Cook, Luis Hernandez, Joseph Jorgeson, Kendall Queen, Cody Summerson, Gabriela Vazquez, and Ava Wallace led by teachers Mr. Jeff Newton and Mrs. Irina Stan are exploring fundamental chiral molecular self-assembly at the atomic scale. Students meet weekly to research, collect data, and analyze as true scientists would. The title of their research project is “Exploring fundamental chiral molecular self-assembly at the atomic-scale: In search of clues to the origins of life on Earth”. The abstract of the research that got their project accepted is as follows.
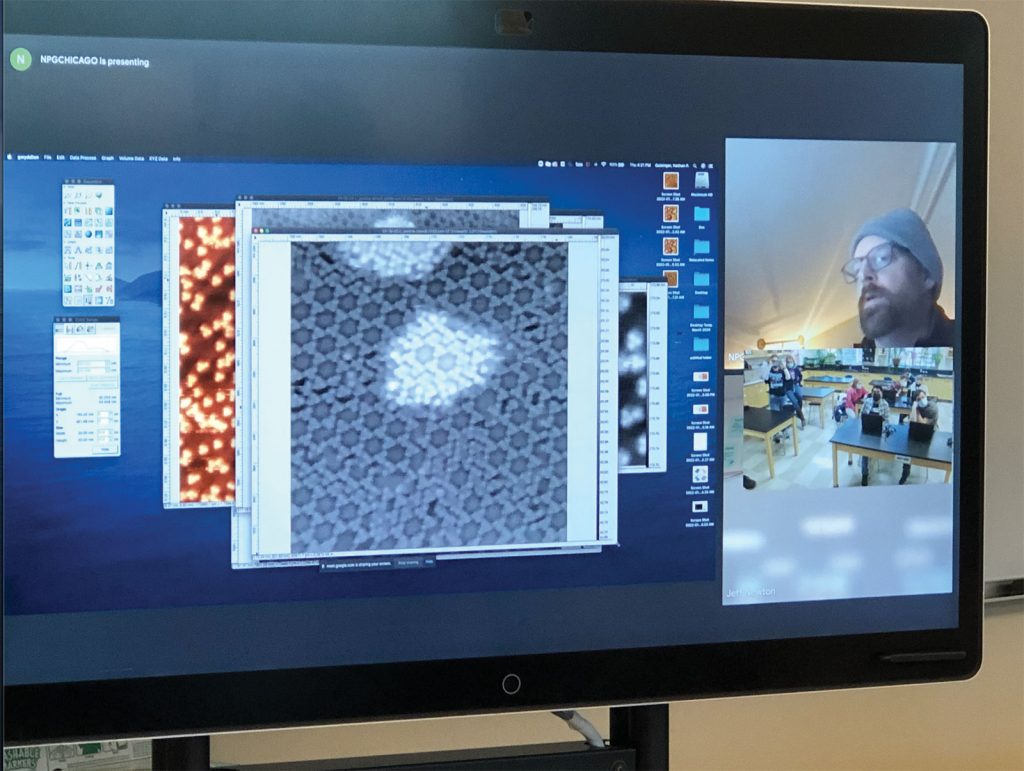
There are more than five hundred amino acids in nature, but only twenty are used in proteins by all living organisms. Amino acids play a central role in cellular metabolism, synthesis of proteins, chemical messengers in communication between cells, and many other biologically important functions. Due to the roles amino acids play in developing the human body, we may be able to use them in drugs to help with bodily functions such as protein creation and tissue repair. The first step toward creating these drugs and medical processes will be to understand how amino acids interact with themselves and other substances. Completing this project will grant us knowledge about how two of the amino acids, histidine and cysteine, self-assemble into structures and how their chirality affects their interactions. Specifically, we will perform fundamental research utilizing an ultrahigh vacuum (UHV) scanning tunneling microscope (STM) to perform fundamental studies on the chirality of amino acid self-assembly on single crystal metal surfaces [1-2]. We will observe chirality in the larger ordered molecular assembly, and we hypothesize that the chirality in L- and D- versions of a particular amino acid will be translated into larger assemblies. We will initially explore the amino acids histidine and cysteine on Cu(111) surface, and we will explore the molecular assembly of homochiral L-, D- amino acids and racemic mixtures (50% L-, 50% D-). The only way to ensure the amino acids self-assemble without interference from outside particles is to perform the experiment in a UHV. Likewise, a STM is the best way to get the most accurate data on the self-assembly of the amino acids. This is why the atomic resolution of the UHV STM is critical for characterization of these surfaces.













